Welcome to CompuCell3D
Congratulations Joel
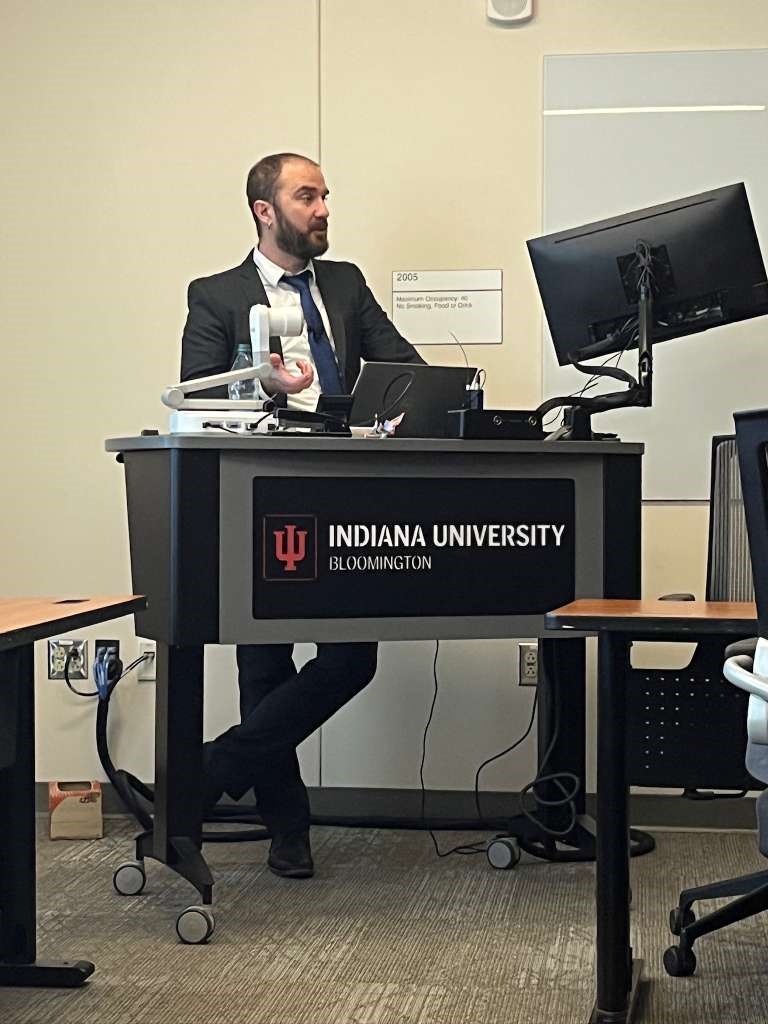
Congratulations to Joel Vanin for successfully defending his CC3D-based PhD thesis, "Mechanistic Virtual Tissue Models as Trustworthy Scientific Instruments". Here's one of his papers: V-Cornea: A computational model of corneal epithelium homeostasis, injury, and recovery |
|
NEW CC3D Version 4.8.0 (March 6, 2026)
We are pleased to announce the new version 4.8.0 of our software CompuCell3D. This release includes many new features and several bug fixes. For more details, please visit our Downloads page: Downloads (opens in new tab with Ctrl/Cmd+Click).
Upcoming Competitions and Workshops
2026 CompuCell3D Multicell Virtual Tissue Modeling Workshop & Hackathon
We are pleased to announce the 21st Annual Multicell Virtual-Tissue Modeling Online Summer School and Hackathon for 2026! This year's workshop will take place virtually between Monday, July 27th and Sunday, August 9th, 2026. We would very much appreciate your help in letting your colleagues and students know about this opportunity. |
Modeling the Invisible: Competition on Forecasting Viral Spread with Limited Data
In this competition workshop, participants will address the challenge of selecting and parameterizing models for complex biological systems. Biological systems present unique hurdles due to their complexity and variability. Even in the face of this complexity, methods to predict the future state of biological systems (and populations) from limited time series data is an ongoing challenge in medical and public heath areas. |
Workshop Dates: Wed. June 3, 2026 - Sat. June 6, 2026 |
Website: Modeling the Invisible Workshop |
Location: Georgia State University, Atlanta GA |
Brochure: Modeling the Invisible Brochure |
Application Deadline: April 22, 2026 |
Contact: invisibleworkshop@gsu.edu |
New Seminars available on Multiscale Modeling with Compucell3D
James Sluka, Indiana University, presentation "Predicting disease with ‘broken’ models, a work in progress" to the IMAG/MSM Viral Pandemics and GLIMPRINT groups |
|
Prof. James Glazier (Indiana U.) and Prof. TJ Sego (U. Florida) presentation "Multiscale Multicellular Modeling: Current Challenges and Future Directions" to the Center for Reproducible Biomedical Modeling. |
|
Prof. James Glazier's COMBINE 2023 Talk: "Multiscale Muticellular Agent-Based Virtual Tissue Simulations: Challenges and Opportunities in Sharable Virtual Tissue Model Specification". |
|
Dr. Juliano Ferrari Gianlupi's COMBINE 2023 Talk: "Breaking Barriers in Multiscale Agent-Based Models: A Path for Cross-Platform Models". |
34 Years!
A thirtieth anniversary! Back on March 16th, 1992, François Graner and James Glazier submitted our very first paper on the Cellular Potts Model/Glazier-Graner-Hogeweg model to Physical Review Letters. We had no idea at that point that the method would still be used today and would be implanted in a dozen different modeling frameworks. As of today, that original paper (which appeared on September 28th, 1992) has been cited more than 1350 times. |
Our Collaboratory Projects
We collaborate extensively with several other computational biology tools groups. In particular:
Tissue Forge
Tissue Forge is a continuation of the Mechanica project. The goal of Tissue Forge is to deliver a modeling and simulation framework that lets users from all relevant backgrounds interactively create, simulate and explore models at biologically relevant length scales. We believe that accessible and interactive modeling and simulation is key to increasing scientific productivity, much like how modeling environments have revolutionized many fields of modern engineering.
Mechanica
 Mechanica is an interactive particle based physics, chemistry and biology simulation environment, with a heavy emphasis on enabling users to model and simulate complex sub-cellular and cellular biological physics problems. Mechanica is part of the Tellurium project.
Mechanica is an interactive particle based physics, chemistry and biology simulation environment, with a heavy emphasis on enabling users to model and simulate complex sub-cellular and cellular biological physics problems. Mechanica is part of the Tellurium project.
Mechanica is designed first and foremost to enable users to work interactively with simulations -- so they can build, and run a simulation in real-time, and interact with that simulation whilst it's running. The goal is to create an SolidWorks type environment where users can create and explore virtual models of soft condensed matter physics, with a emphasis towards biological physics.
- Mechanica is a native compiled C++ shared library with a native and extensive Python API, that's designed to used from an iPython console (or via scripts of course).
Tellurium
 Tellurium allows you to model, simulate, and analyze biochemical systems using a single tool.
Tellurium allows you to model, simulate, and analyze biochemical systems using a single tool.
- Tellurium is a Python Environment for Reproducible Dynamical Modeling of Biological Networks
- Tellurium provides the interfacial code to convert between standard formats and utilize powerful libraries without requiring technical expertise, allowing you to focus on what’s important: building better models.
Tellurium also provides first-class support for exchangeability via COMBINE archives, allowing you to share your models and simulations with other tools.
Invitation to join the IMAG/MSM MULTISCALE MODELING AND VIRAL PANDEMICS Working Group
The invitation to join the IMAG/MSM working group is here.
NEWS: We have viral infection model papers!
"A Multiscale Multicellular Spatiotemporal Model of Local Influenza Infection and Immune Response", Sego, T. J., Mochan, E. D., Ermentrout, G. B., & Glazier, J. A. (2021). Journal of theoretical biology, 532, 110918. Full Text
"Multiscale Model of Antiviral Timing, Potency, and Heterogeneity Effects on an Epithelial Tissue Patch Infected by SARS-CoV-2", Ferrari Gianlupi J, Mapder T, Sego TJ, Sluka JP, Quinney SK, Craig M, Stratford RE Jr, Glazier JA. Viruses. 2022 Mar 14;14(3):605. doi: 10.3390/v14030605. PMID: 35337012; PMCID: PMC8953050. Full Text
NEWS: Our Covid-19 virtual tissue model is available for running on nanoHUB!
You can run our model in your browser, without any installations, at  (here).
(here).
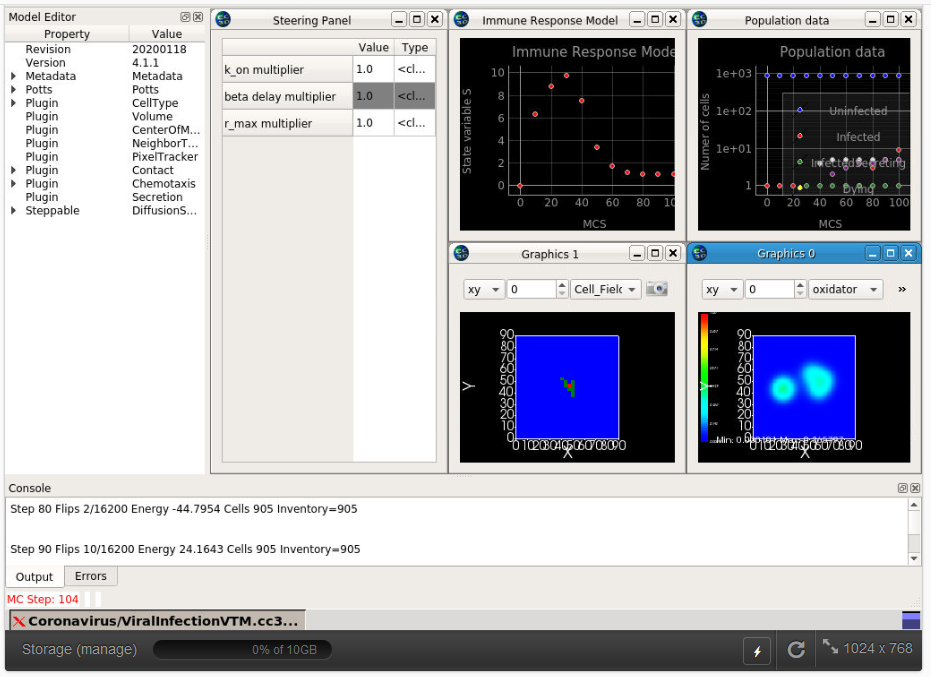
For more info please visit our CC3D on nanoHUB page.
NEWS: CompuCell3D Multiscale, Virtual-Tissue Spatio-Temporal Modeling of Simulations of COVID-19 Infection, Viral Spread and Immune Response and Treatment Regimes
Simulations of tissue-specific effects of primary acute viral infections like COVID-19 are essential for understanding differences in disease outcomes and optimizing therapeutic interventions. In this two-part mini-workshop we present an open-source Python and CC3DML-scripted multiscale model and simulation of an epithelial tissue infected by a virus, a simplified cellular immune response and viral and immune-induced tissue damage and show how you can use it to model basic patterns of infection dynamics and antiviral treatment. Part I presents the model and teaches how to run it and to change model parameters for generating new biologically meaningful simulations. Part II teaches how to extend the model with additional images, graphics and file outputs, additional cell types, diffusive fields, cell behaviors and interactions and improved subcellular and immune-system models.
For more info and scheduling see here. The Part I and Part II videos are now available on YouTube.
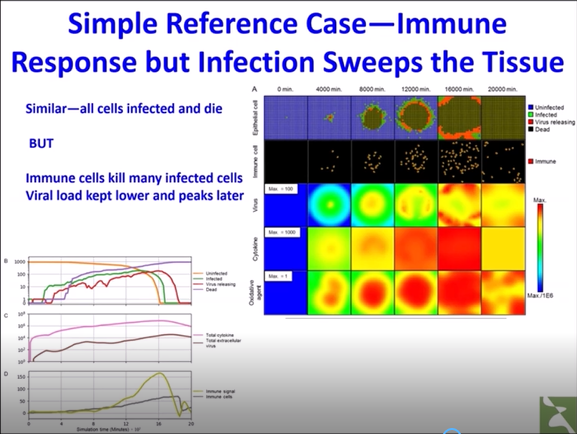
We have prepared a YouTube video describing the model described in our preprint.
Multiscale multicellular modeling of tissue function and disease using CompuCell3D: A simplified computer simulation of acute primary viral infection and immune response in an epithelial tissue
This is a more detailed video presentation hosted by the Pacific Institute for the Mathematical Sciences on the use of CC3D in Covid-19 modeling.
(Click on this image to view the video.)

Call to Contribute: Collaborative Tissue Model Development to Mitigate the Coronavirus Pandemic
The rapidly evolving social and medical impact of COVID-19 requires equally rapid development of tools to predict, and ultimately ameliorate, the future course of the pandemic. Tools are needed to predict patient outcomes, to optimize the deployment of resources by medical providers, and to optimize therapy for infected individuals. Predictive computational simulations of the spread of the disease, and of the underlying biology that makes COVID-19 unique, are both essential. Our aim is to build cooperatively-developed, open-source multiscale model components that can predict various aspects of infection at molecular and tissue scales and inform higher-level models, and a framework to allow others to do so efficiently.
You can run our model online without any installations in https://nanohub.org/tools/cc3dcovid19.
COVID-19 Model Repository: Because of the urgency of the crisis, software development must be done in parallel and not sequentially as in normal scientific model development. As such, we are developing a model repository for the cooperative development of public, open-source molecular- and tissue-scale models and simulations of various infection events and health states associated with COVID-19. To support rapid parallel development, integration, and sharing of models and simulations, we are developing modeling and simulation tools specific to this project, as well as documentation tools and standards for rapid and consistent dissemination of repository contents. Please join us in our efforts to collectively mitigate the ongoing pandemic and future pandemics alike on GitHub. For more information, please contact us at <tjsego AT iu DOT edu>.
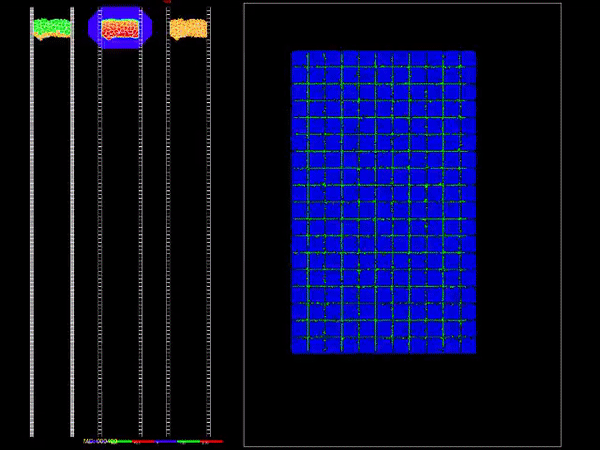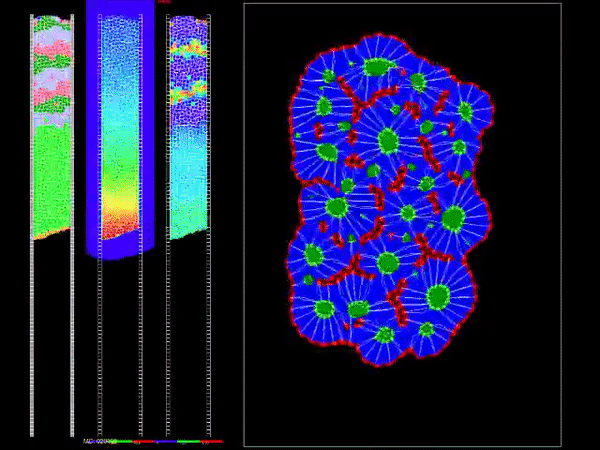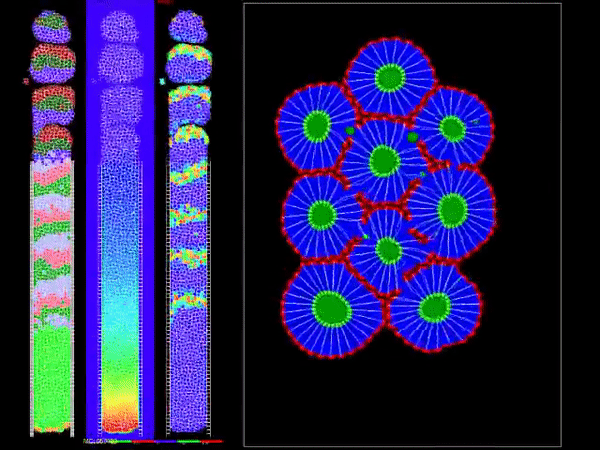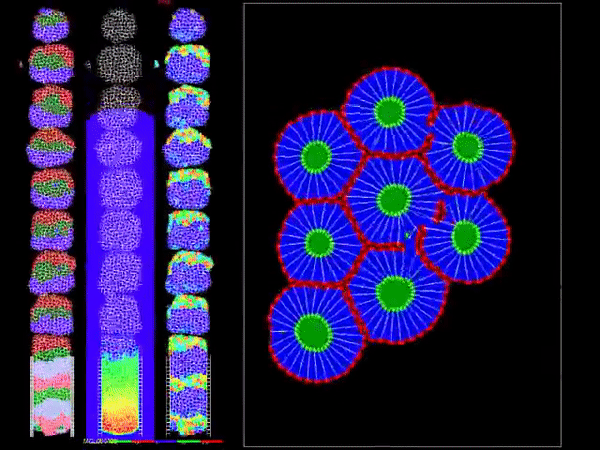
Model of Viral Tissue Infection: In support of parallel model development of tissue infection, the members of the Biocomplexity Institute are developing a multiscale computational model of viral tissue infection using CC3D. The model describes select interactions between generalized epithelial and immune cells and their extracellular environment associated with viral infection and immune response at the cellular and intracellular levels, and in the context of spatiotemporal dynamics. The model is a generic viral infection model, and our hope is to collaboratively develop it into a model of SARS-CoV-2 tissue infection and Covid-19 progression. As such, it is intended to serve as a base model for constructing and implementing more advanced models of targeted cellular- and intracellular-level phenomena in tissue after initial exposure. In its current state, it has not been formally peer-reviewed, and should not be used for patient diagnostics or predicting clinical outcomes. Rather, the model and its implementation can be used to develop and interrogate mechanistic hypotheses about the spread of a virus and how the interplay between viral spreading and immune response determine the outcome of the disease, such as:
- Why does the progression of the disease seem to be dependent on the initial viral exposure level?
- Why is the start time of symptoms and immune response so variable?
- What is the role of cytokine signaling in explaining immune response variability?
Model documentation and simulation files can be found in the project folder of our COVID-19 Model Repository. Please be advised that this is a rapidly evolving project, so model and simulation features are subject to change (even daily). For more information about how to contribute to, modify, or reuse the model and simulation, please contact us at <tjsego AT iu DOT edu>.
For information on recent modeling and repository developments, current development plans, and opportunities to contribute to current modeling and repository projects, please see our project page.
CompuCell3D on ...

CompuCell3D is available for online use on nanoHUB. Some CompuCell3D simulations are already available as demonstrations. See the CC3D on nanoHUB page for more info.
About CompuCell3D
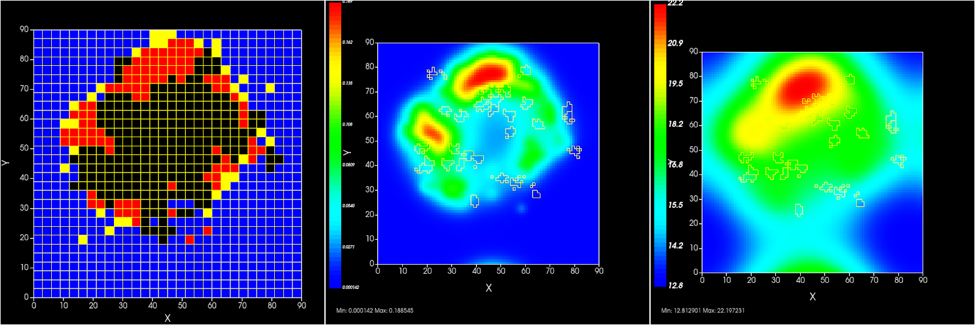
CompuCell3D is a flexible scriptable modeling environment, which allows the rapid construction of sharable Virtual Tissue in silico simulations of a wide variety of multi-scale, multi-cellular problems including angiogenesis, bacterial colonies, cancer, developmental biology, evolution, the immune system, tissue engineering, toxicology and even non-cellular soft materials. CompuCell3D models have been used to solve basic biological problems, to develop medical therapies, to assess modes of action of toxicants and to design engineered tissues. CompuCell3D's intuitive interface makes Virtual Tissue modeling accessible to users without extensive software development or programming experience.
It uses Cellular Potts Model to model cell behavior. Below is a preview of what CC3D can do:
Support
This project is currently funded by generous support from the U.S. National Science Foundation (NSF) grant NSF-1720625, “Network for Computational Nanotechnology - Engineered nanoBIO Node” and
National Institute of Health, National Institute of Biomedical Imaging and Bioengineering (NIBIB) Grant U24EB028887, "Dissemination of libRoadRunner and CompuCell3D".
Past support for this project includes:
U.S. National Institutes of Health (NIH) grant NIH - R01 GM122424, “Competitive Renewal of Development and Improvement of the Tissue Simulation Toolkit”. Information on previous support is listed on our Support page.
CompuCell3D is led by James A. Glazier (Indiana Univ) in collaboration with Dr. David Umulis (Purdue Univ). Many people contribute, or have contributed, to CC3D including the original lead developer Maciej Swat.
New CC3D forum
To report any bugs, or to ask questions, please visit our Q&A page: Reddit CompuCell3D.
Note: In April 2019 our old user forum at CC3D - AllAnswered was discontinued. The entries in the old AllAnswered forum are available at CC3D AllAnswered archive.html. The text of the questions and answers are there but the linked files are not. If you need one of the linked files please ask for it on the Reddit forum.
Want to contribute to CompuCell3D development?
We highly encourage fellow researchers to contribute to CompuCell3D code. If you have developed some useful plugin or functionality on CompuCell3D and would like share it with rest of us. Please commit the code and open a pull request on CompuCell3D GitHub repository. We would love to see it in next version of CompuCell3D.
How to cite CompuCell3D
Multi-Scale Modeling of Tissues Using CompuCell3D – M. Swat, Gilberto L. Thomas, Julio M. Belmonte, A. Shirinifard, D.Hmeljak, J. A. Glazier, Computational Methods in Cell Biology, Methods in Cell Biology 110: 325-366 (2012). PMID:22482955 PMCID: PMC3612985 DOI: doi.org/10.1016/b978-0-12-388403-9.00013-8
New Papers using CompuCell3D Simulations have been published:
For a more complete list see our Publication Page.
Adhesion-driven invasion: Disentangling the interplay between cell-cell and cell-matrix interactions in cancer cell migration. Quirine J.S. Braat, Klara Beslmüller, Cornelis Storm, Erik H.J. Danen, Liesbeth M.C. Janssen. Biophysical Journal, 2026, ISSN 0006-3495, https://doi.org/10.1016/j.bpj.2026.02.025.
Injury-specific muscle regeneration: a computational blueprint for cellular and cytokine drivers. Haase, Megan, Tien Comlekoglu, Alexa Petrucciani, T. J. Sego, Shayn Peirce, and Silvia Blemker. Journal of The Royal Society Interface 23, no. 236 (2026).
A model of tuberculosis progression using CompuCell3D. J. W. G. Doran, C. F. Rowlatt, G. G. Powathil, R. Bowness and C. A. Yates, 2026, arXiv, https://arxiv.org/abs/2602.24258.
Start small: A model for tissue-wide planar cell polarity without morphogens.Thayambath, Abhisha, and Julio M. Belmonte. PLOS Computational Biology 22, no. 2 (2026): e1013938.
Simple cellular Potts model of scratch assays on healthy and keloid fibroblasts driven by contact inhibition of locomotion. Urcun, Stéphane, Yasmin El Mahi, Raluca Eftimie, Zélie Dirand, Gwenaël Rolin, and Stéphane PA Bordas. Journal of Theoretical Biology, Volume 622, 7 April 2026, 112365.
Glycation‐Driven Impairment of Cytoskeletal Homeostasis and Viability Disables Glyoxalase‐Low Mesothelia From Resisting Cancer Colonization. Mishra, Satyarthi, Kottpalli Vidhipriya, Anchita Gopikrishnan, Purba Sarkar, Sudiksha Mishra, Chigicherla VS Prasanna, P. S. Hari, Aruna Korlimarla, Prosenjit Sen, and Ramray Bhat. Journal of Cellular Physiology 241, no. 1 (2026): e70123.
Polyclonal and single clonal patient-derived organoid models of Barrett oesophagus and oesophageal adenocarcinoma establish a platform for the analysis of heterogeneity in disease progression and therapy response. Jacobson, Daniel H., Dylan P. McClurg, Emily Black, Charlotte Cassie, Tik Shing Cheung, Hannah Coles, Ginny Devonshire et al. bioRxiv (2026): 2026-01.
Cortical tension links curvature to tissue growth in the cellular Potts model. Fastabend, Kai Lennard, Cécile M. Bidan, John WC Dunlop, and Philip Kollmannsberger. Physical Review E 113, no. 2 (2026): 024403.
A Multiscale Computational Model of Endothelial-Immune Cell Interactions Regulated by Dynamic Wall Shear Stress.Zhang YY, Wang YT, Li YJ, Qiu Y, Chen D, Qin KR. Int J Numer Method Biomed Eng. 2026 Jan;42(1):e70137. doi: 10.1002/cnm.70137. PMID: 41521423.
 CompuCell3D
CompuCell3D