= Building CompuCell3D on Red Hat Enterprise Linux 6 and CentOS 6= Building CompuCell3D from source on Unix/Linux systems is fairly straightforward once all of it's dependencies have been satisfied. The following commands should build and install CC3D on RHEL 6 systems.
Prerequisites
Hardware
CompuCell3D may build and run with less capable hardware, but has been tested with the following:
- 1024MB RAM
- Hardware 3D Graphics Acceleration (most modern graphics cards)
Build Tools and Dependencies
we will assume that you have already installed Software Developer Workstation Package Group from Red Hat 6 distribution. Typically when you install RHEL 6 you have a choice of the package groups and Software Developer Workstation is one of such groups. If you talk to your sys-admin they can certainly help you install this package set. RHEL ships with Python 2.6 by default and CC3D runs best with Python 2.7. Therefore we will build CC3D based on Miniconda Python distribution from Continuum Analytics.
Using miniconda package manager we will install
- pyqt4
- numpy
- scipy
- vtk
This will save us a lot of time compared to the manual compilation of those packages.
CompuCell3D requires also the following packages to be present:
- swig
- cmake
- QScintilla (with python bindings)
- pcre-devel
We will install those packages either using yum package manager or will compile them from source.
Let's start with the easy part first -swig and cmake installation. Open a terminal and become root by typing :
su -
enter your root password and type the following:
yum install cmake-gui swig pcre-devel
Now let's install miniconda - here I will choose to perform a local installation i.e. I will install it in my home directory. First let's get appropriate package - by visiting http://conda.pydata.org/miniconda.html
and choosing Python 2.7 64-bit installer. Navigate to your download directory and make the downloaded script executable :
chmod +x Miniconda2-latest-Linux-x86_64.sh
After that, run the installation by typing from your download directory
./Miniconda2-latest-Linux-x86_64.sh
Towards the end of the installation you will be asked if you want the installer script to add path to miniconda Python distribution to the PATH environment variable. I recommend you say yes.
If you change your mind you can always open .bashrc script and remove path to miniconda. Adding path to miniconda python will make it your default Python and this is a good thing on a system that ships with obsolete version of Python. Additionally installing additional Python packages using miniconda toolset is super easy. So let's do it- I will show you how to install pyqt, numpy vtk, and scipy using miniconda. I assume that if you type
conda
you will get conda usage help. If not then for the reminder of presented installation steps you will need to specify full path to the conda program.
Let's install the required dependencies now:
conda install pyqt numpy scipy vtk
Answer yes if conda asks whether to install the dependencies, wait few minutes and you will have most of cc3d dependencies needed to run CC3D on your machine.
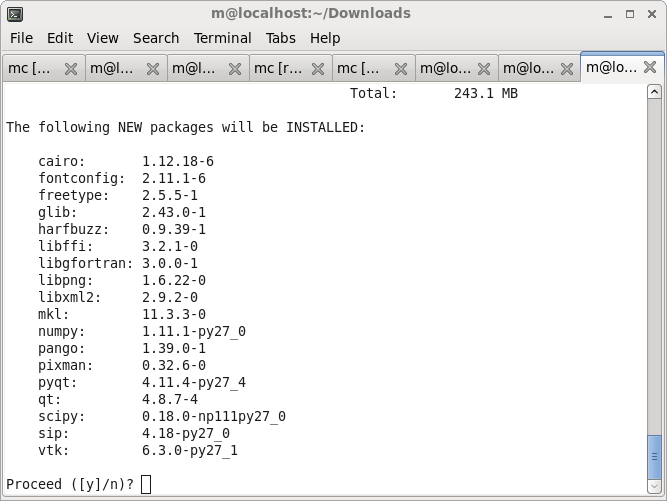
To check if everything went OK open a terminal and type
python
followed by
from PyQt4 import QtCore
The output should look as shown below:
Now we will need to compile two packages
- QScintilla (with Python bindings) - needed by CC3D player and Twedit++
PyQwt 5.2.0 (needed for plotting capabnilities within Player)
Let's start with QScintilla. First let's navigate to QScintilla download page - https://www.riverbankcomputing.com/software/qscintilla/download
We will get source code tarball for Linux. The installation instructions are in http://pyqt.sourceforge.net/Docs/QScintilla2/
and I suggest you follow them. But first let's make sure that you are picking the correct installation of the Qt. You want qt bundled with miniconda. To make sure this is the case type
qmake --version
The output should show that indeed your shell sees qt version bundled with miniconda:
If this is not the case simply prepend path to miniconda's bin directory to your PATH environment variable.
If everything works as follows you should be able to follow the installation instructions for QScintilla"
cd Qt4Qt5 qmake qscintilla.pro make make install
After QScintilla is build we will build python bindings for it. Assuming we are in the Qscintilla source main directory we would type
cd Python python configure.py make make install
At this point we should have QScintilla and Python bindings to it installed on our machine
Now let's look at the installation of plotting library PyQwt-5.2.0
You get it from https://sourceforge.net/projects/pyqwt/files/pyqwt5/PyQwt-5.2.0/PyQwt-5.2.0.tar.gz/download
Unpack it and go to the unpacked PyQwt folder
You should be able to follow the installation instructions shown here - http://pyqwt.sourceforge.net/doc5/installation.html
Here is my console window showing basis steps of PyQwt installation:
One thing you have to be aware of here. There is a small bug in the PyQwt source code. Fortunately it is very easy to fix. Here we go - navigate to PyQwt installation folder located inside miniconda folder. In my case since miniconda was installed in
/home/m/miniconda2
the folder I am looking for is
/home/m/miniconda2/lib/python2.7/site-packages/PyQt4/Qwt5
We will fix anynumpy.py script as follows - we will replace original code (I am shogin only first part of the file- look for lines following if name =='numpy': )
# import either NumPy, or numarray, or Numeric
for name in ('numpy', 'numarray', 'Numeric'):
failed = False
try:
eval(compile('from %s import *' % name, 'eval', 'exec'))
if name == 'numpy':
from numpy.oldnumeric.compat import *
Float = float
UInt8 = uint8
except ImportError:
failed = True
if not failed:
breakwith
for name in ('numpy', 'numarray', 'Numeric'):
failed = False
try:
eval(compile('from %s import *' % name, 'eval', 'exec'))
if name == 'numpy':
pass
#from numpy.oldnumeric.compat import *
#Float = float
#UInt8 = uint8
except ImportError:
failed = True
if not failed:
breakIf you prefer downloading the fixed file , you can do it by clicking here
At this point we are done with the dependencies
Source Code
Once the dependencies have been satisfied make directory where you want to store source code, in my case it is in /home/m/CC3D_GIT
mkdir /home/m/CC3D_GIT cd /home/m/CC3D_GIT
Once the directory has been created, obtain the source code from our GIT repository using the following command:
git clone https://github.com/CompuCell3D/CompuCell3D.git .
This will clone CC3D Git repository into current directory (remember about the . at the end of last command - it is important)
Build Configuration: Starting CMake
CompuCell3D is configured using the CMake build system. The following command starts the CMake GUI:
cmake-gui
That will start the build system. Having miniconda is in your PATH environment variable will help locate the dependencies . Click Browse Source... and select the CompuCell3D source directory from the CompuCell3D GIT directory we have created above - in my case the CompuCell3D source directory is located in /home/m/CC3D_GIT/CompuCell3D
In addition to specifying source directory we also specify the location of the build directory i.e. a directory where compilation files will be stored. in my case it is /home/m/CC3D_GIT_build/CompuCell3D
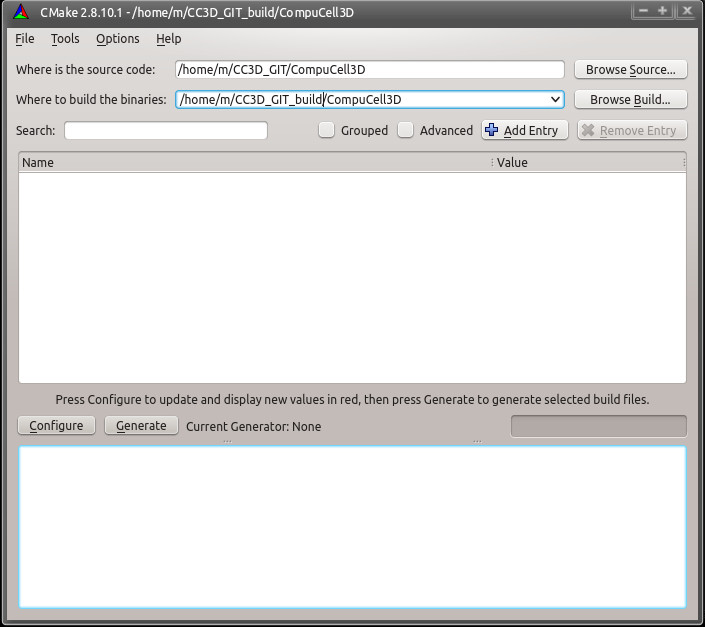
We are ready to click Configure. A dialog box asking to create the build directory will appear:
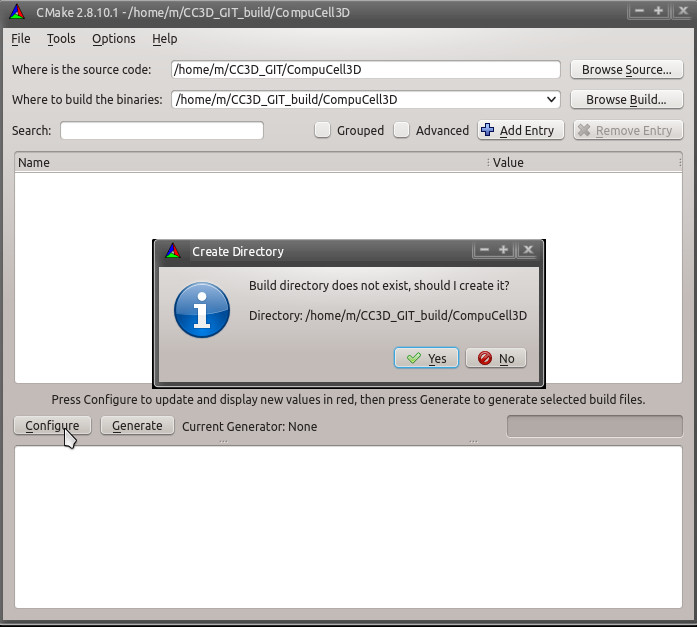
and then one asking about the build system:
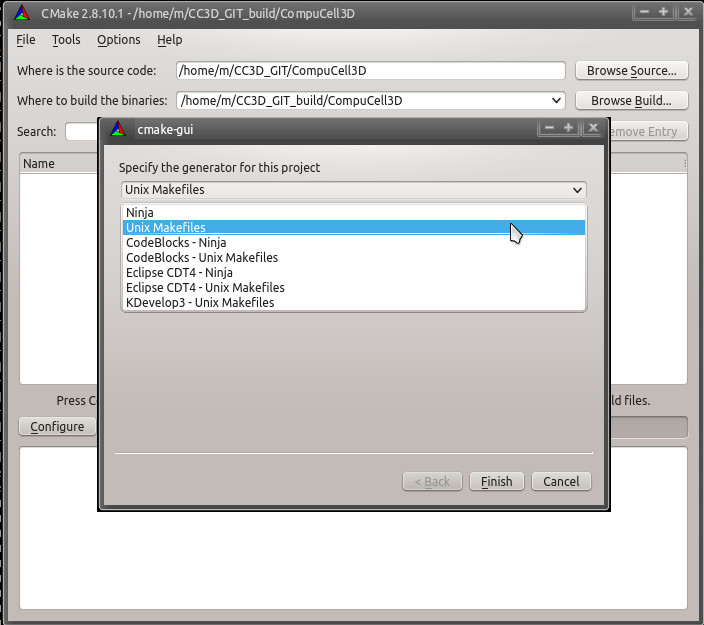
Select Unix Makefiles for the generator, select Use default native compilers and then click Finish. CMake will begin the configuration process. CMake will attempt to locate all of the dependencies installed above.
After the initial configuration has completed select Grouped and Advanced to make entering configuration values easier.
Build Configuration: CMAKE
In CMAKE confirm that CMake has located all the build tools. In the CMAKE_BUILD_TYPE field you may enter Debug, RelWithDebInfo or Release to specify the type of the binary you want to have - if you are developing extra modules compiling in the Debug or RelWithDebInfo can be helpful. By default Compucell3D build type is set to Release:
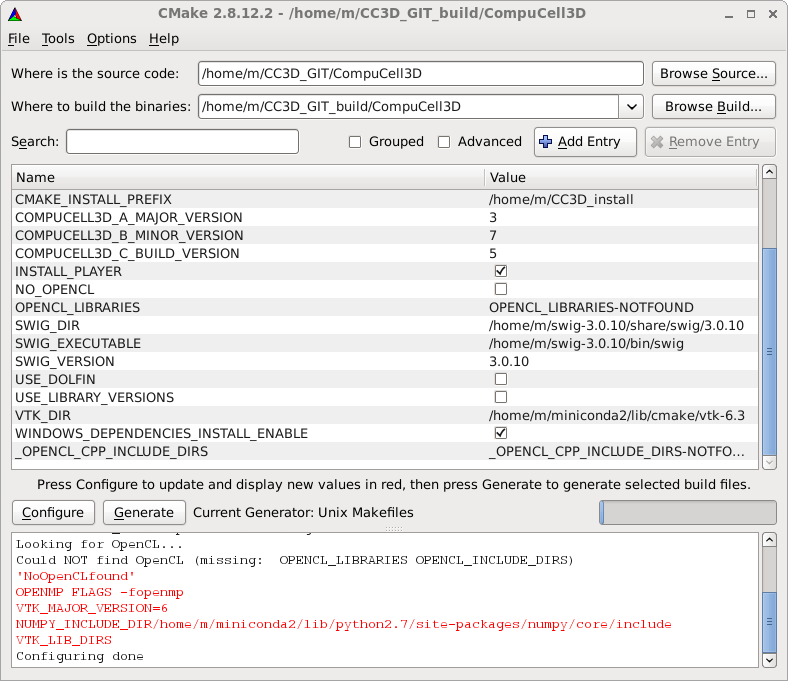
In the CMAKE_INSTALL_PREFIX field enter the directory you would like to install CompuCell3D into - this is important. We recommend that unless necessary you should install CC3D into local directory - in my case it is /home/m/CC3D_install, .
Click on Configure to have all the values updated.
IMPORTANT CMake will try to locate all the dependencies and Python is one of the dependencies. Alas, CMake script that locates Python distribution (Python executable, library and header directory) mixes different Python distributions if you have more than one Python distribution on your system. Therefore it is important to fix the appropriate paths in the cmak-gui before proceeding further. To do this. Check Advanced box, and in the search line next to the box type python and you will see the following Python dependencies listed:
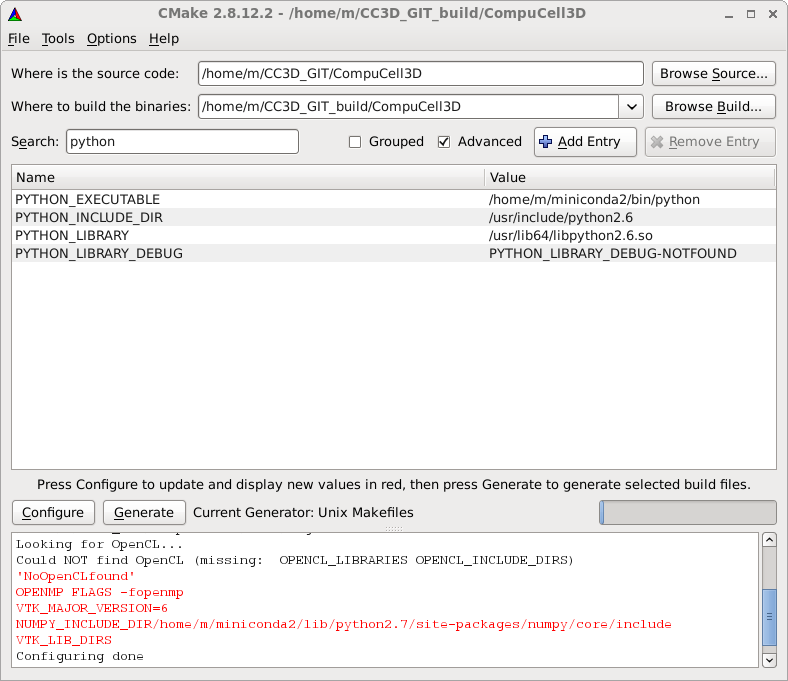
As you can see Python executable comes from miniconda distribution while library and header files come from system-default Python (which is obsolete 2.6 edition). We need to fix it, specifying paths to the directory containing Python.h file and the path to the python library. In my case those are /home/m/miniconda2/include/python2.7 and /home/m/miniconda2/lib/libpython2.7.so respectively - see screenshot below
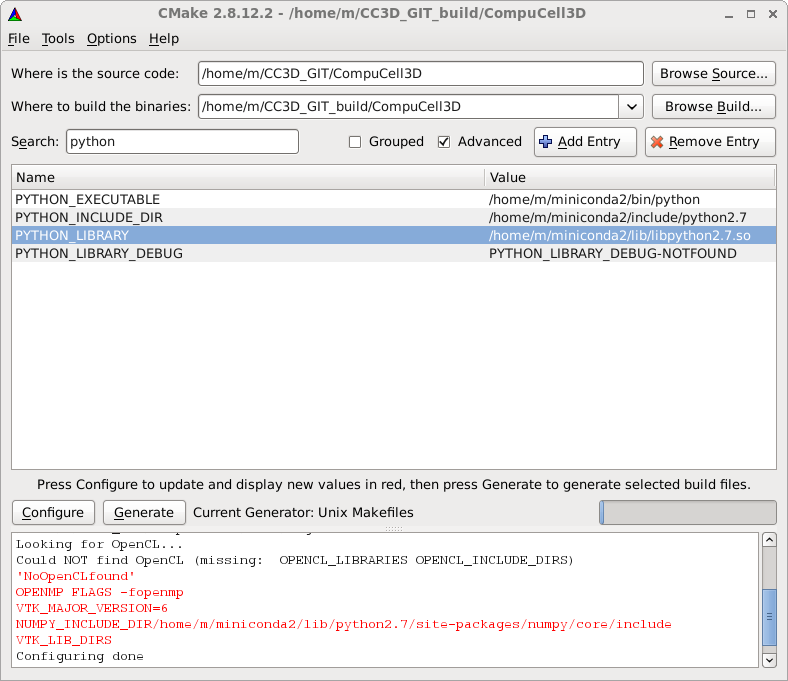
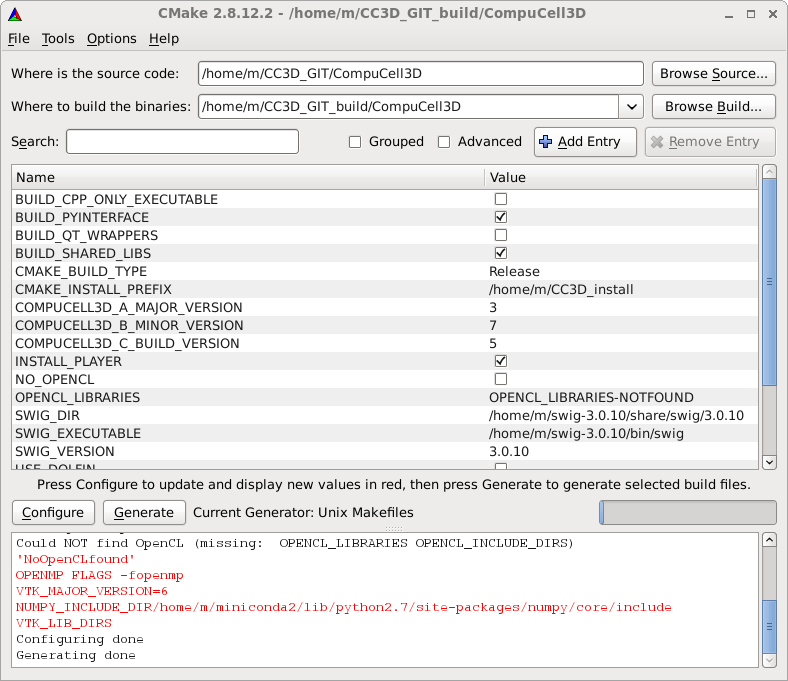
After this fix we can move further - click on Configure and then click on Generate to generate the Makefiles, ignore any warnings. Exit CMake
Building
To begin compiling CompuCell3D enter the build directory specified in the Where to build the binaries field of CMake earlier and use the following command:
In my case I type:
cd /home/m/CC3D_GIT_build/CompuCell3D make
After the build begins you may see screen like this one:
Installing
Once compiling has completed CompuCell3D can be installed into the directory specified in the CMAKE_INSTALL_PREFIX field earlier by issuing the following command:
make install
Running
After CompuCell3D has finished installing you can start CompuCell3D by entering the installation directory (in my case it is in /home/m/CC3D_install):
cd /home/m/CC3D_install
and issuing the following command:
./compucell3d.sh
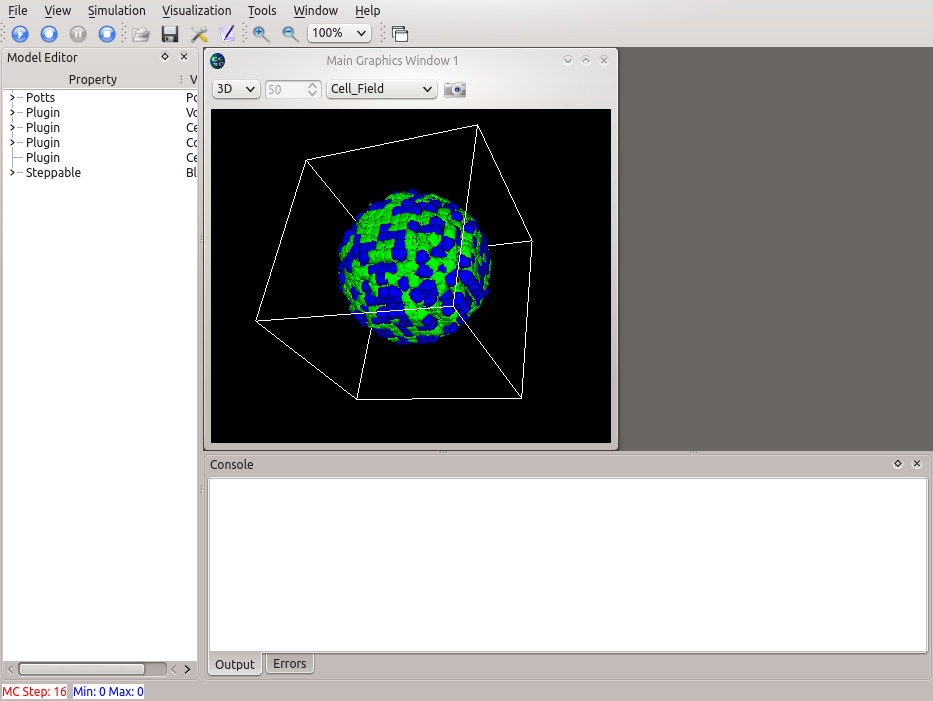
 CompuCell3D
CompuCell3D